Introduction
Infertility is described as an inability to conceive after one year of unprotected intercourse, with a total prevalence of 9% [1]. Primary and secondary infertility has been defined as childlessness and failure to conceive in a woman who has already had one child or more, respectively [2]. Infertility can occur in both genders and in different ways. Genetic, anatomical, immunological and endocrinological ab-normalities can lead to infertility. The factors affecting male infertility, include quality, motility, sperm counts and ejaculatory dysfunctions [3].
In 20% of infertile couples, there is at least one male fertility defect, and can reach over 40% [4]. The main causes of male infertility are varicocele (37%), semen disorders (10%), testicular insufficiency (9%), obstruction (6%), cryptorchidism (6%), and other abnormalities (7%). Additionally, about 25% of male infertility cases are unexplained, being referred to as idiopathic infertility [5]. Many studies have examined the genetic causes of male infertility, but only 15% of infertility cases have been detected [6]. Consequently, there is still a need for a better understanding of male infertility, and we must consider other approaches to identify its causes. Epigenetic is one of the promising approaches that can partly explain the causes of idiopathic cases. Therefore, studying the epigenetic basis of male infertility may be crucial in managing an infertile patient.
Discussion
The role of the epigenetic factors in male infertility
The epigenetic modifications are alterations in phenotype caused by mechanisms that do not change the DNA sequence [7]. These alterations are excluded in sperm for two reasons. First, the occurrence of the elimination of epigenetic marks in primordial germ cells (PGCs). Second, the occurrence of the genomic condensation and reorganization in male germ cell nuclei [8]. The most common epigenetic modifications include DNA methylation, histone modifications, transition from canonical histones to protamines and non-coding RNAs (ncRNAs) [9]. The process of adding a methyl group to the 5' position on the cytosine pyrimidine ring, called as DNA methylation, occurs in hot spot regions (CpG islands) [10]. Abnormalities in this process can affect significant processes, including spermatogenesis, and may lead to male infertility [11, 12]. Also, for replacement of histones with protamines, several proteins are involved, including P1 and P2. The mutations affecting these proteins can lead to sperm abnormalities and infertility [13]. The two main types of ncRNAs are infrastructural and regulatory. The major function of the infrastructural ncRNAs is in translation and splicing, and include rRNAs, tRNAs, and small nuclear RNAs. However, regulatory ncRNAs are often implicated in the epigenetic processes and include small non-coding RNAs (sncRNAs) [14, 15]. The sncRNAs act as the gene expression regulators in various cellular processes. The most important sncRNAs are miRNA, siRNA, piRNA and lncRNA, their differences of which are presented in table 1 [16, 17].
The piRNAs as a non-coding RNA
For the first time in 2006, a novel class of small non-coding RNAs was isolated from the mouse testis and the Drosophila germ cells called P-element induced wimpy testis in drosophila (PIWI)-interacting RNAs (piRNAs) [18, 19]. The length of the piRNAs is about 26-33 nucleotides. About 86% of piRNAs have a uracil deflection at the 5' end and play a crucial role in spermatogenesis [20]. According to the origin of piRNAs, they can be divided into three classes: a-piRNAs originated from retrotransposons, b-piRNAs originated from mRNA, and c-piRNAs originated from lncRNAs (long noncoding RNAs) [21]. The most common piRNAs are derived from retrotransposons and are linked to Ago3 (Argonaute), PIWI and AUB (Aubergine) proteins [22, 23]. The main function of piRNAs in germ cells is the suppression of transposon activity [24].
Biogenesis of piRNA
The main sites of the piRNAs distribution are the animal testis spermatogonial cells, ovarian oocytes and Drosophila follicle cells (somatic cells).
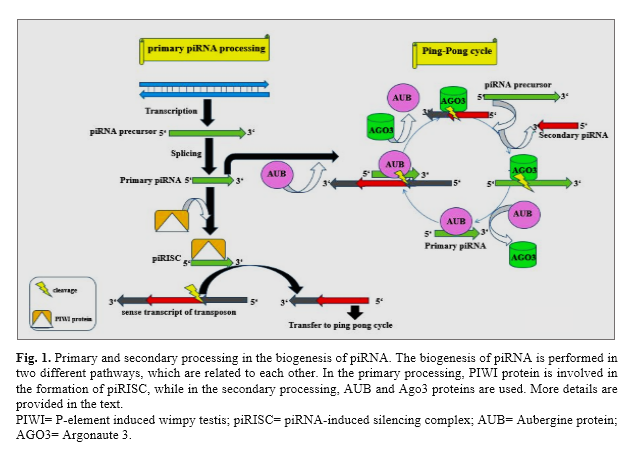
There are two main pathways for the piRNA biogenesis: In the germ cells, the AUB-dependent pathway (secondary processing) is active while in the somatic cells, the only pathway for producing piRNAs is the PIWI-dependent pathway (primary processing) [25]. The primary antisense transcript of piRNAs is preferably attached to PIWI protein and produces the piRNA-induced silencing complex (piRISC). The piRISC cleaves the sense transcript of transposons at positions 10 and 11 and generates the 5' end of Ago3-associated piRNA. In the secondary processing known as the 'ping–pong' cycle, the AUB and Ago3 proteins are involved [26]. Like PIWI, AUB protein plays a pivotal role in the formation of the piRISCs and produces the 5′ ends of piRNAs that are associated with Ago3 [27]. Also, the Ago3 complex has two crucial functions: First, Ago3 produces secondary piRNAs by targeting Ago3-associated piRNAs. Second, Ago3 produces the 5′ end of the antisense piRNAs through the cleavage of antisense piRNA precursors and then are loaded onto AUB (Figure 1) [28]. The HEN1 protein mediates 2′‑O-methylation of the 3′ end of piRNA. In the mice, there are two PIWI proteins including MILI and MIWI2 which produce piRNAs through processing of transposable elements (TEs). This occurs in cytoplasmic granules known as pi-bodies and piP-bodies [29].
Table 1. The types of small non-coding RNAs (sncRNAs) and their distinctive features
| ncRNAs |
Length |
Main cell type |
Dicer-dependent |
Estimated number |
Main functions |
| miRNAa |
19-25 |
Male germ cell |
Yes |
Less than 1000 |
Translational repression |
| siRNAb |
19-29 |
Oocyte |
Yes |
More than 10,000 |
mRNA cleavage |
| piRNAc |
26-31 |
Various |
No |
More than 10,000 |
Transposon silencing |
| lncRNAd |
>200 |
Various |
No |
More than 4,000 |
Chromatin remodeling and transcriptional regulation |
a: microRNA; b: small interfering RNA; c: PIWI interacting RNA; d: long non-coding RNA
The role of piRNAs in male infertility
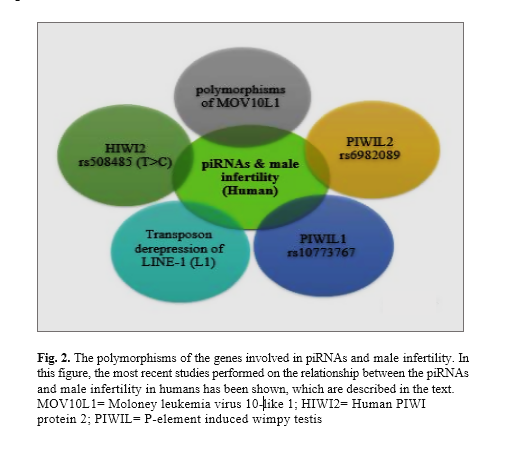
The piRNAs can play different roles in biological processes, including sex determin-ation, gene silencing, epigenetic regulation, and cancer. One of the most important functions of the PIWI-piRNA complex is to protect the genome of gametes against the transposon invasion and is performed through silencing their transcripts [27]. Consequently, piRNAs are usually useful for the human genome, but the aberrant expression of genes involved in the biogenesis can lead to changes in the genome and the development of the various disorders. Male infertility is one of the disorders associated with piRNAs. In figure 2, the most important research performed on male infertility and piRNAs is summarized: Moloney leukemia virus 10-like 1 (MOV10L1) is a gene associated with the biogenesis of piRNA that plays a role in the primary and secondary processing [30]. The MOV10L1 can contribute to the primary piRNAs for binding to the PIWI proteins. Some studies confirm that several polymorphisms of this gene show a remarkable increase in infertile men [31]. In human, the relationship between four PIWI proteins (HIWI, HILI, HIWI2 and PIWIL3) and male infertility is shown. In 2010 and 2017, investigations on the Chinese and Iranian population with non-obstructive azoospermia independently revealed a relationship between HIWI2 rs508485 (T>C) and non-obstructive azoospermia and can be considered as a risk factor for male infertility [32, 33]. Furthermore, transposons are transposable elements that use the host genome to survive and amplify [34]. The PIWI-piRNA complexes act as genomic guardians of the gametes against transposon invasion and target the transposons by silencing transcripts. LINE1 is a known retrotransposon. Studies on patients with cryptorchidism have revealed that infertility in patients is due to alterations in the PIWI-pathway and derepression of transposable elements [35]. Therefore, these studies show that piRNAs may play a crucial role in male infertility.
The potential role of piRNAs as a diagnostic biomarker for male infertility
According to the World Health Organization (WHO), diagnosis of male infertility is based on the semen parameters which include the following: motility, sperm concentration, seminal volume, pH and morphology [36]. Some studies have indicated that the sperm analysis cannot be used accurately to detect between fertile and infertile men [37]. Hence, identification of non-invasive seminal biomarkers can solve this problem. Cell-free RNAs and sncRNAs are critical as non-invasive biomarkers in controlling pregnancy and detecting reproductive-related disorders [38, 39]. In 2015, Hong and colleagues, determined five piRNAs by examining seminal plasma samples of infertile patients, which can be used as diagnostic biomarkers for the detection of infertile men [40]. Also, another study on patients with idiopathic male infertility who had experienced the first intracytoplasmic sperm injection course demonstrated that there is a relationship between spermatozoa piRNA levels (piR-31704 and piR-39888) and sperm concentration [41]. As a result, these piRNAs are important factors in the fertilization process.
The piRNAs and DNA methylation
DNA methylation, as an epigenetic marker, is associated with many disorders. In germ cells, methylation is implicated in the TE silencing, genomic imprinting, and DNA compaction. Investigations have revealed that the loss of CpG methylation on LINE1 in germ cells is related to the dysfunction of the piRNA pathway. Therefore, it is thought that this pathway plays an important role in the identification of active LINE1 elements and the guidance of the methyltransferase complexes for the methylation of active elements [42]. Moreover, earlier studies have shown that abnormal methylation occurs in men with low sperm quality [43]. The mutation in the genes involved in the piRNAs processing can be associated with human spermatogenic disorders.
A recent study on peripheral blood samples of infertile men has indicated that rs10773767 and rs6982089 were two single nucleotide polymorphisms in PIWIL1 and PIWIL2, respectively. These polymorphisms are allele-specific methylation-sensitive [44]. Thus, methylation changes in these genes are associated with spermatogenic disorders. Therefore, TDRD1 (a Tudor-domain-containing protein) protein is involved in the regulation of spermatogenesis by interacting with MIWI [45]. The loss of this interaction can activate the transposons [46].
Some variants of this gene are associated with a risk of spermatogenesis defects and infertility [47, 48]. Additionally, due to the relationship between the modified pattern of TEs methylation and male infertility [48, 49], these alterations may be caused by the dysfunction of the piRNAs pathway. Therefore, it is suggested that studying the methylation patterns in the pathways of piRNAs processing can help to better understand the etiology of male infertility.
The targeting of piRNAs as novel therapies
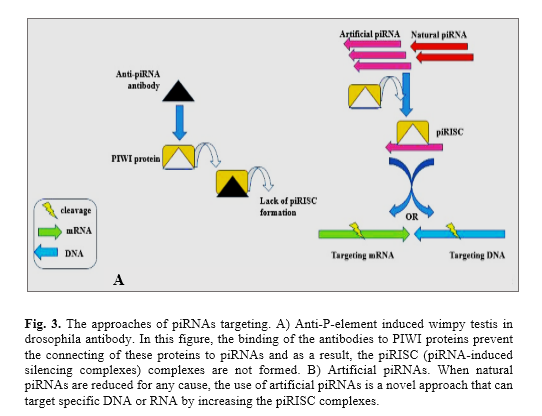
The sncRNAs, as one of the new therapies, can be used for different disorders. Significant advances have been made in the field of clinical applications of miRNAs, such as the use of Miravirsen (an oligonucleotide with 15-nucleotide) in the treatment of the human immunodeficiency virus infections [50], miRNAs replacement and inhibition techniques in the treatment of many cancers [51, 52]. Also, based on the function of piRNAs and PIWI proteins, there are two approaches to change the piRNAs expression: the antibodies can be useful against PIWI proteins at post-translational levels while artificial piRNAs are a good option for both transcriptional and post-translational approaches (Fig. 3) [53]. The anti-PIWI antibodies prevent the formation of the piRISCs, hence, the piRNA expression can be changed. Moreover, decreased or absent the expression of genes associated with piRNAs may lead to an increased expression level of transposons. In this case, the use of artificial piRNAs can be a useful approach. Additionally, in germ cells, transposons may alter DNA methylation and induce methylation through artificial piRNAs, that can lead to silencing of the gene expression [54]. Therefore, they are important in evaluating inherited epigenetic alterations. However, these new approaches are in the early stages and need more extensive research.
 Conclusions
Conclusions
One of the benefits of understanding the epigenetic abnormalities is that epigenetic modifications, unlike genetic mutations, can be modified using specific drugs. Hence, with
the complete understanding of these changes, treatment of epigenetic-related diseases can be achieved. The sncRNAs are the most common epigenetic regulators and their role is identified in many disorders. Among sncRNAs, piRNAs play a key function in spermatogenesis and are a candidate for further investigations on male infertility. The studies presented in this review revealed that investigating the role of piRNAs in male infertility can be useful for many causes: first, determining a non-invasive biomarker for early detection of male infertility, second, discovering the causes of idiopathic male infertility. Also, piRNAs can be used to identify types of infertile patients. For example, piR-30198 is one of the piRNAs used for this purpose. This biomarker is able to distinguish between the two disorders related to male infertility, namely, azoospermia and asthenozoospermia [40].
Conflict of Interest
The authors declare to have no conflict of interest.
Acknowledgments
The authors appreciate the valuable contributions of the experts in the Research and Clinical Center for Infertility of Yazd, especially in the molecular and cytogenetic laboratories.
References











































