Introduction
It has been estimated that gastric mucosa of more than half of the world’s population is infected with Helicobacter pylori (H. pylori), ranging from 25-50% in the developed countries to more than even 80% in the developing world [1, 2]. The prevalence of infection in 10% of children in the industrialized countries and more than 50% in the developing countries shows that H. pylori infection is usually found in childhood and in most cases remains in the gastric mucosa for a lifetime unless treated [1-3]. Chronic inflammation in the mucosal epithelium due to persistent H. pylori infection is associated with a variety of clinical outcomes such as gastritis and gastric ulcers in 10–20% of the infected individuals, which are the most common type of gastric diseases related to H. pylori [3]. Additionally, it is well established that H. pylori infection, classified as category I carcinogen, plays a crucial role in the development of gastric cancer, and 1-2% of infected subjects develop this type of cancer [4]. In addition to bacterial and environmental factors that affect the individuals’ susceptibility to H. pylori infection, host factors are also important [5]. Several polymorphisms in the genes encoding inflammatory proteins, cytokines, growth factors, chemokines and another immune response genes have been identified to be associated with the susceptibility to H. pylori infection [6, 7]. Cells involved in the innate immunity including mucosal epithelial cells and myeloid cells, macrophages and dendritic cells, are responsible for the initial response to H. pylori components during infection which is mediated through the pattern recognition receptors (PRRs) such as toll-like receptors (TLRs). TLRs recognize preserved microbial structures, pathogen-dependent molecular patterns (PAMPs), such as flagellin, lipopoly-saccharide (LPS) and peptidoglycan located on the surface of the bacteria, fungi, protozoa, and viruses. TLRs also recognize endogenous ligands such as damage-associated molecular patterns (DAMPs) e.g. heat shock proteins which are spontaneously released during cell stress [8]. The family of TLRs includes 13 members in mammalians that are either located on the surface of the cell, such as TLRs 1, 2, 4, 5 , 6 and 10, or are located on the endoplasmic reticulum membrane or on the endosomal/ lysosomal membrane such as TLRs 3, 7, 8 and 9 [8-10]. The interaction of TLRs, expressed in most tissues and components of the envelope and nucleic acids of the infectious pathogen, results in the activation of intracellular signaling pathways that lead to inflammatory responses and activation of immune responses [8, 9]. Recently, several investigations have shown the role of genetic variations, including single nucleotide polymorphisms (SNPs), in the members of the TLR family in susceptibility to various infections, especially bacterial infections [9].
The purpose of this systematic review and meta-analysis was to evaluate the most prevalent TLR polymorphisms in H. pylori-positive subjects and the association of these polymorphisms with the susceptibility to H. pylori infections.
Materials and Methods
Literature search strategy
Three authors independently searched the electronic databases including PubMed, Scopus, Google Scholar and ISI web of knowledge to collect all case-control studies evaluating polymorphisms in the TLR 1 to 13 genes and their association with susceptibility to H. pylori infection. The last search was performed on January 15, 2019 using Medical Subject Headings (MeSH) terms including toll-like receptors, TLRs, polymorphisms, single-nucleotide polymorphisms, SNPs, mutations, variations and H. pylori infection. Hand searching of reference lists of the selected studies was performed to find any missed potential relevant article.
Study selection criteria
Eligible articles were selected based on the following criteria: 1) papers published in the English language, 2) case-control studies; case samples were infected individuals with H. pylori (H. pylori positive) and controls were H. pylori uninfected (H. pylori negative) and 3) articles investigating polymorphisms in the TLR 1 to 13 genes and their association with susceptibility to H. pylori infection. Articles with the following characteristics did not meet our inclusion criteria and were excluded from the meta-analysis: 1) articles evaluating the association between polymorphisms in the TLRs genes and susceptibility to other bacterial infection or other diseases, 2) articles evaluating other gene polymorphisms or other host genetic factors, 3) articles not comprising enough information on the prevalence of polymorphisms in the TLRs genes, genotypes and alleles frequencies in H. pylori positive and negative individuals and 4) articles evaluating polymorphisms in the TLRs genes only in the H. pylori infected or H. pylori uninfected individuals.
Data extraction
Data from each study were extracted and tabulated in Table 1. Collected characteristics were as follows: 1) country of origin, 2) year of publication 3) ethnicity, 4) genotyping method, 5) number of H. pylori positive subjects, 6) number of H. pylori negative subjects, 7) type of disease, 8) type of polymorphism, 9) H. pylori detection method and 10) references.
Statistical analysis
We performed all statistical analyses using Comprehensive Meta-Analysis (CMA) software version 2.2 (Biostat, Englewood, NJ, USA). Using allelic and genotypic models, the association between TLRs polymorphisms and susceptibility to H. pylori infection was measured by ratio (ORs) with 95% confidence intervals (95% CIs). When the p value was less than 0.05, association was statistically significant. Additionally, the frequency of polymorphisms in the TLRs genotypes/alleles among H. pylori infection positive case and control subjects was expressed as percentage (%). Fixed-effects model was used to pooled data when there was no heterogeneity and Cochrane Q test was statistically significant (I2=0–25%, P<0.05); however, in large heterogeneity (I2=25–100 %, P<0.05), random effects model was used [11]. In the current meta-analysis, publication bias was evaluated using funnel plots.
Results
Study and populations characteristics
After comprehensive literature search in the electronic databases including PubMed, Scopus, Google Scholar and ISI web of knowledge until January 15, 2019, a total of 3157 articles were collected. As shown in Figure 1, 18 qualified studies were included for final analysis after meeting the inclusion criteria, including 5 studies from Caucasian ethnicity, 3 studies from Iranian ethnicity, 2 studies from Chinese ethnicity, 2 studies from Japanese ethnicity, 2 studies from Thai ethnicity, 1 study from Malaysian ethnicity, 1 study from Indian ethnicity, 1 studies from Kashmiri ethnicity and 1 study from Brazilian ethnicity. The main characteristics of included studies in the meta-analysis are shown in table 1. In the present study, included studies were reported from China, Japan, Thailand, Malaysia, India, Iran, Germany, Scotland, Lithuania, Latvia and Brazil. Main TLRs genes polymorphisms in included studies which are more common among H. pylori infection positive individuals were shown in the different ethnicities (Table 1). Polymerase chain reaction (PCR)-based methods were the most frequent techniques used for determining the TLRs genotypes/alleles. Additionally, various studies have used different detection methods to identify H. pylori positive subjects. Patients had different H. pylori-related diseases including gastric cancer, non-ulcer dyspepsia, peptic ulcer disease, gastric atrophy, mucosa-associated lymphoid tissue lymphoma and gastritis. Table 2 displays distribution of genotypes and alleles frequencies of TLRs genes polymorphisms in H. pylori positive and negative individuals. Additionally, Table 3 reveals association between TLRs genes polymorphisms, genotypes/ alleles, and H. pylori infection.
Association between TLR1 polymorphisms and susceptibility to H. pylori infections
In the present meta-analysis, we evaluated possible associations between TLR1 rs4833095 and TLR1 rs5743618 polymorphisms and susceptibility to H. pylori infections (Table 2).
The TLR1 rs4833095 variants were CC, CT and TT that occurred in 896 (45.8%), 687 (35.2%) and 370 (19%) of H. pylori infection-positive cases and in 416 (29%), 871 (60.5%) and 151 (10.5%) H. pylori infection-negative controls. Frequency of mutant genotypes, CT and TT, were 1057 (54.1%) and 1022 (71%) in H. pylori infection-positive and -negative individuals, respectively. In the Chinese, Thai, and Malaysian populations, genotype frequency of variant genotypes, CT and TT, were 859 (44%), 164 (8.3%) and 34 (1.7%), respectively, in H. pylori infection-positive individuals and 639 (44.4%), 364 (25.3%) and 19 (1.3%), in H. pylori infection-negative individuals, respectively. Overall, in the genotypic model, there was no significant association between TLR1 rs4833095 polymorphism and susceptibility to H. pylori infections (Table 3). For the TLR1 rs5743618 polymorphism, genotypes, II, IS and SS, were 14 (6.2%), 72 (31.4%) and 143 (62.4%) in H. pylori infection-positive cases and 4 (7.5%), 17 (32%) and 32 (60.5%) in H. pylori infection-negative controls. Additionally, in Caucasian population, frequency of mutant genotypes, IS and SS, were 215 (93.8%) and 49 (92.4%) in H. pylori infection-positive and -negative individuals, respectively. Such as TLR1 rs4833095 polymorphism, there was no significant association between TLR1 rs5743618 polymorphism and susceptibility to H. pylori infections (Table 3).
Association between TLR2 polymorphisms with susceptibility to H. pylori infections
Several studies have reported association between six TLR2 SNPs (-196 to -174 (ins→del), rs3804099 (T→C), rs3804100 (T→C), +2251 (G→A), rs121917864 (Arg677Trp), and rs5743708 (Arg753Gln)) with susceptibility to H. pylori infections.
For TLR2 -196 to -174 SNP, genotype and allele frequencies were as follows: ins/ins 541 (46.2%), ins/del 504 (43.2%), del/del 124 (10.6%), ins 345 (70.5%), and del 145 (29.5%) in H. pylori infection-positive individuals and ins/ins 376 (46%), ins/del 355 (43.6%) and del/del 85 (10.4%), ins 198 (72.7%), del 74 (27.3%) in H. pylori infection-negative individuals. Frequency of variant allele, del, and variant genotypes, ins/del and del/del, in different populations are illustrated in table 2. Overall, no association was found between TLR2 -196 to -174 polymorphism with susceptibility to H. pylori infections in genotypic and allelic models (Table 3). For TLR2 rs3804099 SNP, TT, TC and CC genotypes frequency were 126 (61.7%), 27 (13.3%) and 51 (25%) in H. pylori infection-positive individuals and 131 (66.8%), 41 (21%) and 24 (12.2%) in H. pylori infection-negative individuals. In addition, genotype frequency of mutant heterozygote and homozygote, TC and CC, were 78 (38.2%) and 65 (33.1%) in H. pylori infection-positive and -negative individuals, respectively. Individuals with the CC mutant homozygote genotype for TLR2 rs3804099 SNP had a significantly increased risk of H. pylori infection (Table 3 and Fig. 2). For TLR2 rs3804100 SNP, TT, TC and CC genotypes frequency were 147 (72%), 49 (24%), and 8 (4%) in H. pylori infection-positive individuals and 149 (76%), 43 (22%), and 4 (2%) in H. pylori infection-negative individuals. Genotype frequency of mutant heterozygote and homozygote, TC and CC, were 57 (28%) and 47 (24%) in H. pylori infection-positive and -negative individuals, respectively. The results indicated that the TLR2 rs3804100 TC and CC genotypes variant failed to have any association with increased risk of H. pylori infection in Thai population (Table 3). For TLR2 +2251 (G→A) SNP, GG, GA and AA genotypes frequency were 225 (99.1%), 2 (0.9%) and 0 (0%) in H. pylori infection-positive individuals and 252 (98.4%), 4 (1.6%) and 0 (0%) in H. pylori infection-negative individuals. Genotype frequency of mutant genotypes, GA and AA, were 2 (9%) and 4 (1.6%) in H. pylori infection-positive and -negative individuals, respectively. In Brazilian population, the TLR2 +2251 GG and GA genotypes variant did not show any association with increased risk of H. pylori infection (Table 3).
Association between TLR4 polymorphisms and susceptibility to H. pylori infections
In the current meta-analysis, association between five TLR4 SNPs (rs4986790, rs4986791, rs11536889, rs10759932, and rs1927914) and susceptibility to H. pylori infections was evaluated. For TLR4 rs4986790 SNP, frequency of genotypes amounted to AA 1104 (84.5%), AG 192 (14.7%) and GG 9 (0.8 %) in H. pylori infection-positive cases and 981 (85.5%), 157 (13.6%) and 9 (0.9 %) in H. pylori infection-negative controls. Frequency of mutant genotypes, AG and GG, were 201 (15.4%) and 166 (14.4%) in H. pylori infection-positive and -negative individuals, respectively. Additionally, frequency of mutant allele, G, was 45 (17%) and 47 (8.5%) in H. pylori infection-positive and -negative individuals, respectively. Overall, no association was found between TLR4 rs4986790 polymorphism with susceptibility to H. pylori infections in genotypic model; however, there was a significant association in allelic model (Table 3 and Figure 2). For TLR4 rs4986791 SNP, CC, CT and TT genotypes frequency were 131 (72.4%), 41 (22.6%), and 9 (5%) in H. pylori infection-positive individuals and 223 (87.1%), 27 (10.5%), and 6 (2.4%) in H. pylori infection-negative individuals. Frequency of variant allele, T, and variant genotypes, CT and TT, were as follows: 57 (21.4%) and 50 (27.6%) in H. pylori infection-positive individuals, and 36 (6.5%) and 33 (12.8%) in H. pylori infection-negative individuals. Both mutant homozygote genotype, TT, and mutant allele, T, have shown highly significant association with susceptibility to H. pylori infections (Table 3 and Fig. 2).
For TLR4 rs11536889 SNP, GG, GC and CC genotypes frequency were 1016 (59.4%), 587 (34.2%), and 109 (6.4%) in H. pylori infection-positive individuals and 431 (59.8%), 242 (33.6%), and 47 (6.6%) in H. pylori infection-negative individuals. Also, G and C alleles frequency were 984 (85%) and 170 (15%) in H. pylori infection-positive individuals and 614 (85%) and 112 (15%) in H. pylori infection-negative individuals. Frequency of variant allele, C, and variant genotypes, GC and CC, were as follows: 170 (14.7%) and 696 (40.6%) in H. pylori infection-positive individuals, and 112 (15.4%) and 289 (40.1%) in H. pylori infection-negative individuals. Overall, no association was found between TLR4 rs11536889 polymorphism with susceptibility to H. pylori infections in genotypic and allelic models (Table 3). For TLR4 rs10759932 SNP, TT, TC and CC genotypes frequency leveled at 153 (75%), 37 (18.2%), and 14 (6.8%) in H. pylori infection-positive individuals and 139 (71%), 53 (27%), and 4 (2%) in H. pylori infection-negative individuals. We found a significant association between TLR4 rs10759932 CC mutant genotype with susceptibility to H. pylori infections (Table 3 and Fig. 2). Our results revealed a significant association between TLR4 rs1927914 T mutant allele with susceptibility to H. pylori infections (Table 3 and Figure 2).
Association between TLR5 polymorphisms and susceptibility to H. pylori infections
Among four TLR5 SNPs (rs5744174, rs5744168, rs1640827 and rs17163737) which were evaluated in terms of their susceptibility to H. pylori infections, only one polymorphism, TLR5 rs5744168, yielded enough information. Genotype frequency of CC, CT, and TT were 206 (91.6%), 19 (8.4%), 0 (0%) in H. pylori infection-positive individuals and 239 (93.7%), 16 (6.3%), and 0 (0%) in H. pylori infection-negative individuals. Frequency of variant genotypes, CT and TT, were 19 (8.4%) in H. pylori infection-positive individuals, and 16 (6.2%) in H. pylori infection-negative individuals. Overall, no association was found between TLR5 rs5744168 polymorphism and susceptibility to H. pylori infections in genotypic model (Table 3).
Association between TLR9 polymorphisms with susceptibility to H. pylori infections
We evaluated the role of four TLR9 SNPs (rs352140, rs34399053, rs150459369 and rs5743836) in susceptibility to H. pylori infections. For TLR9 rs352140 SNP, genotype frequency of CC, CT, and TT were 25 (32.5%), 46 (59.7%), and 6 (7.8%) in H. pylori infection-positive individuals and 100 (43.4%), 98 (42.6%), and 32 (14%) in H. pylori infection-negative individuals. In addition, C and T alleles frequencies were 96 (62.3%) and 58 (37.7%), in H. pylori infection-positive individuals and 298 (64.8%) and 162 (35.2%) in H. pylori infection-negative individuals. Frequency of variant allele, T, and variant genotypes, CT and TT reached as follows: 58 (37.7%) and 52 (67.5%) in H. pylori infection-positive individuals, and 162 (35.2%) and 130 (56.5%) in H. pylori infection-negative individuals. Our results demonstrated a significant association between TLR9 rs352140 CT mutant genotype with susceptibility to H. pylori infections (Table 3 and Fig. 2).
For TLR9 rs34399053 SNP, genotype frequency of CC, CT, and TT were 26 (33.7%), 51 (66.3%), and 0 (0%) in H. pylori infection-positive individuals and 79 (34.3%), 150 (65.2%), and 1 (0.5 %) in H. pylori infection-negative individuals. Frequency of variant allele, T, and variant genotypes, CT and TT, were as follows: 51 (33.2%) and 51 (66.3%) in H. pylori infection-positive individuals, and 152 (33%) and 151 (65.6%) in H. pylori infection-negative individuals. On balance, no association was found between TLR9 rs34399053 polymorphism with susceptibility to H. pylori infections in allelic and genotypic models (Table 3). For TLR9 rs150459369 SNP, CC, CT, and TT genotypes frequency were 77 (100%), 0 (0%) and 0 (0%) in H. pylori infection-positive individuals and 220 (95.6%), 10 (4.4%) and 0 (%) in H. pylori infection-negative individuals. In addition, C and T alleles frequency were 154 (100%) and 0 (0 %) in H. pylori infection-positive individuals and 450 (97.8%) and 10 (2.2%) in H. pylori infection-negative individuals. Frequency of variant allele, T, and variant genotypes, CT and TT, were as follows: 0 (0%) and 0 (0%) in H. pylori infection-positive individuals, and 10 (2.2%) and 10 (4.4%) in H. pylori infection-negative individuals. By and lange, no association was found between TLR9 rs150459369 polymorphism with susceptibility to H. pylori infections in allelic and genotypic models (Table 3). For TLR9 rs5743836 SNP, TT, TC and CC genotypes frequencies were 77 (65.8%), 40 (34.2%), and 0 (0%) in H. pylori infection-positive individuals and 35 (68.6%), 15 (29.5%), and 1 (1.9%) in H. pylori infection-negative individuals. Overall, no association was detected between TLR9 rs5743836 polymorphism and susceptibility to H. pylori infections in allelic and genotypic models (Table 3).
Association between TLR10 polymorphisms with susceptibility to H. pylori infections
The role of two polymorphisms, TLR10 rs10004195 and TLR10 rs4129009, in susceptibility to H. pylori infections were determined. For TLR10 rs10004195 SNP, AA, AT and TT genotypes frequency were 581 (33.3%), 740 (42.3%) and 426 (24.4%) in H. pylori infection-positive individuals and 339 (27.5%), 627 (51%) and 266 (21.5%) in H. pylori infection-negative individuals. Additionally, frequency of variant genotypes, AT and TT, were as follows: 1166 (66.7%) and 893 (72.4%) in H. pylori infection-positive and -negative individuals, respectively. All in all, no association was found between TLR10 rs10004195 polymorphism with susceptibility to H. pylori infections in genotypic model (Table 3). Frequency of variant genotypes, TC and CC, were as follows: 990 (66.1%) and 667 (65.5%) in H. pylori infection-positive and -negative individuals, respectively. Totally, no association was found between TLR10 rs4129009 polymorphism with susceptibility to H. pylori infections in genotypic model (Table 3).
Finally, no study was found in terms of evaluating association between TLR3, 6-8, 11-13 polymorphisms and susceptibility to H. pylori infections.
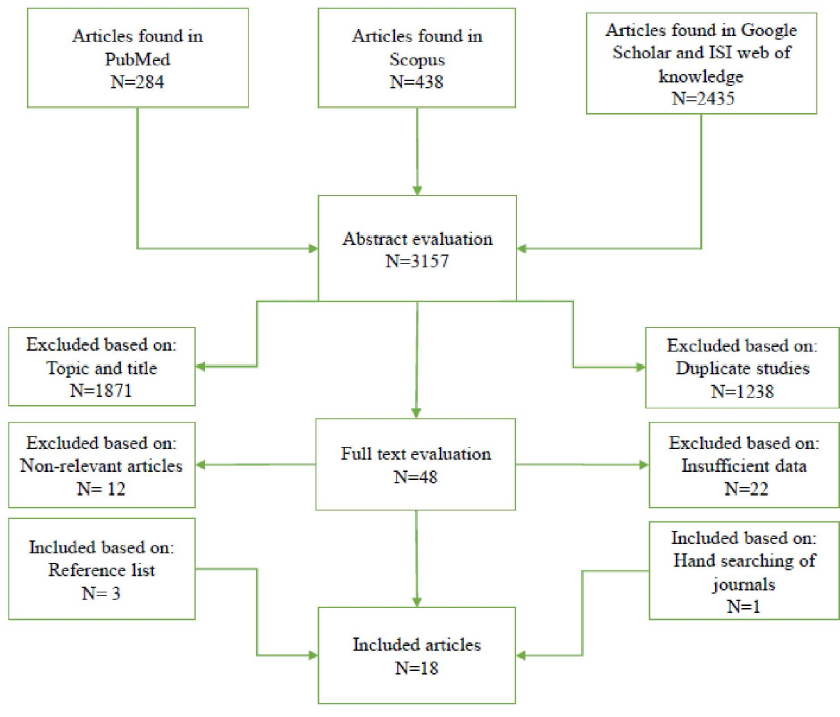
Fig. 1. Flowchart of eligible studies selection process in the meta-analysis
Fig. 2. Forest plot of the meta-analysis on the association between TLR2 rs3804099, TLR4 rs4986790, TLR4 rs4986791, TLR4 rs10759932, TLR4 rs1927914, and TLR9 rs352140 and susceptibility to H. pylori infections.
Table 1. The main data of included studies in the meta-analysis
| Country |
Genotyping method |
H. pylori
positive
(N) |
H. pylori
negative
(N) |
Total
(N) |
Disease type |
Polymorphism |
H. pylori detection method |
Ref. |
| China |
Real-time PCR |
1511 |
1042 |
2553 |
GA
DYS
IM
SG |
TLR1 rs4833095
TLR10 rs10004195
TLR10 rs4129009 |
13C-UBT |
12 |
| China |
Real-time PCR
Mass spectrometry |
190 |
94 |
284 |
GC |
TLR2 -196 to -174 del
TLR4 rs11536889 |
Serum anti-H. pylori
IgG antibody |
13 |
| Japan |
PCR |
937 |
699 |
1636 |
GA |
TLR2 -196 to -174 del |
Serum anti-H. pylori
IgG antibody |
14 |
| Japan |
PCR-CTPP |
1191 |
401 |
1592 |
GA |
TLR4 rs11536889 |
Serum anti-
H. pylori
IgG
antibody |
15 |
| Thailand |
Real-time PCR |
204 |
196 |
400 |
GC
Gastritis
|
TLR1 rs4833095
TLR2 rs3804099
TLR2 rs3804100
TLR4 rs10759932
TLR10 rs10004195 |
Histopathological examination
Real-time PCR |
16 |
| Thailand |
Real-time PCR |
204 |
196 |
400 |
Gastritis |
TLR1 rs4833095 |
Histopathological examination |
17 |
| Malaysia |
Real-time PCR |
62 |
33 |
95 |
GC
NUD
PUD |
TLR1 rs4833095
TLR10 rs10004195 |
RUT Culture Histopathological examination |
18 |
| India |
PCR-RFLP
ARMS-PCR |
77 |
230 |
307 |
PUD |
TLR4 rs1927914
TLR4 rs4986790
TLR4 rs4986791
TLR9 rs352140
TLR9 rs34399053
TLR9 rs150459369
|
RUT |
19 |
| India |
PCR-RFLP |
104 |
26 |
130 |
GC |
TLR4 rs4986790
TLR4 rs4986791
|
PCR (glmM) |
20 |
| Iran |
ASPCR
PCR-RFLP
|
55 |
45 |
100 |
PUD |
TLR2 -196 to -174 ins/del
TLR2 rs121917864
TLR2 rs5743708 |
RUT
Serum anti-H. pylori IgG
antibody
Histopathological examination
PCR (glmM) |
21 |
| Iran |
PCR–CTPP
PCR-RFLP |
56 |
44 |
100 |
PUD |
TLR4 rs11536889
TLR4 rs4986790
TLR4 rs4986791
|
RUT
Histopathological examination
Serum anti-H. pylori
IgG antibody |
22 |
| Iran |
PCR-RFLP |
195 |
241 |
436 |
Gastritis |
TLR4 rs4986790 |
RUT
PCR (16s rRNA and glmM)
*Histological examination |
23 |
| Scotland |
Applied Biosystems 5 nuclease SNP genotyping
assay |
117 |
51 |
168 |
GA |
TLR9 rs5743836 |
14C-UBT Serology
RUT
Culture
Histopathological examination |
24 |
| Scotland |
PCR-RFLP
Real-time PCR |
103 |
46 |
149 |
GC |
TLR4 rs4986790 |
14C-UBT Serology
RUT
Culture
Histopathological examination |
25 |
| Germany |
Fluorescence-labeled hybridization FRET probes |
229 |
53 |
548 |
GC
HRG |
TLR1 rs5743618 |
Serum anti-H. pylori
IgG antibody |
26 |
| Germany |
Real-time PCR |
594 |
358 |
952 |
MALT lymphoma |
TLR4 rs4986790 |
RUT
and
Histopathological examination |
27 |
Germany
Lithuania Latvia |
PCR-RFLP |
203 |
97 |
|
GC
GA |
TLR4 rs11536889 |
Serum anti-H. pylori
IgG antibody |
28 |
| Brazil |
PCR-RFLP |
232 |
254 |
486 |
Duodenal ulcer
Gastritis |
TLR2 (+2251)
TLR4 rs4986790
TLR5 rs5744168 |
13C-UBT Culture
RUT
PCR (urea) |
29 |
MALDI-TOF= Matrix assisted laser desorption ionization time-of-flight; ELISA= Enzyme-linked immunosorbent assay; PCR-RFLP= Polymerase chain reaction-restriction fragment length polymorphism; PCR-CTPP= Polymerase chain reaction with confronting two-pair primers, ASPCR= Allele-specific polymerase chain reaction; HPLC= High-performance liquid chromatography;
13C- and
14C-UBT=
13C- and
14C-urea breath test; RUT= Rapid urease test; GC= Gastric cancer, NUD= Non-ulcer dyspepsia; PUD= Peptic ulcer disease; GA= Gastric atrophy, MALT= Mucosa-associated lymphoid tissue lymphoma; HRG= High-risk gastritis, DYS=; Dysplasia; IM= intestinal metaplasia; SG= Superficial gastritis; NA= Not available.
*Histopathological examination: H & E and Giemsa staining.
Table 2. Distribution of genotype and allele frequencies of polymorphisms in the TLRs genes in H. pylori positive and negative individuals
TLR |
Variant |
SNP
number |
H. pylori infection
positive
(N) |
H. pylori infection
negative
(N) |
Ref. |
| Genotype/Allele |
Genotype/Allele |
TLR1
|
Ser248Asn
(C→T) |
rs4833095 |
CC |
CT |
TT |
C |
T |
CC |
CT |
TT |
C |
T |
|
| 629 |
660 |
199 |
NA |
NA |
379 |
499 |
140 |
NA |
NA |
12 |
| 122 |
4 |
78 |
NA |
NA |
14 |
182 |
0 |
NA |
NA |
16 |
| 23 |
19 |
15 |
NA |
NA |
9 |
8 |
11 |
NA |
NA |
18 |
| 122 |
4 |
78 |
NA |
NA |
14 |
182 |
0 |
NA |
NA |
17 |
Ile602Ser
(I→S) |
rs5743618 |
II |
IS |
SS |
I |
S |
II |
IS |
SS |
I |
S |
|
| 14 |
72 |
143 |
NA |
NA |
4 |
17 |
32 |
NA |
NA |
26 |
TLR2
|
-196 to -174
(ins → del) |
|
ins/
ins |
ins/
del |
del/
del |
ins |
del |
ins/
ins |
ins/
del |
del/
del |
ins |
del |
|
| 418 |
414 |
105 |
NA |
NA |
304 |
316 |
79 |
NA |
NA |
14 |
| 81 |
90 |
19 |
252 |
128 |
46 |
39 |
6 |
131 |
51 |
13 |
| 42 |
NA |
NA |
93 |
17 |
26 |
NA |
NA |
67 |
23 |
21 |
| T→C |
rs3804099 |
TT |
TC |
CC |
T |
C |
TT |
TC |
CC |
T |
C |
|
| 126 |
27 |
51 |
NA |
NA |
131 |
41 |
24 |
NA |
NA |
16 |
T→C |
rs3804100 |
TT |
TC |
CC |
T |
C |
TT |
TC |
CC |
T |
C |
|
| 147 |
49 |
8 |
NA |
NA |
149 |
43 |
4 |
NA |
NA |
16 |
+2251
(G→A) |
|
GG |
GA |
AA |
G |
A |
GG |
GA |
AA |
G |
A |
|
| 225 |
2 |
0 |
NA |
NA |
252 |
4 |
0 |
NA |
NA |
29 |
Arg677Trp
(C→T) |
rs121917864 |
CC |
CT |
TT |
C |
T |
CC |
CT |
TT |
C |
T |
|
| 4 |
NA |
NA |
58 |
52 |
3 |
NA |
NA |
47 |
43 |
21 |
Arg753Gln
(G→A) |
rs5743708 |
GG |
GA |
AA |
G |
A |
GG |
GA |
AA |
G |
A |
|
| 50 |
NA |
NA |
102 |
8 |
44 |
NA |
NA |
89 |
1 |
21 |
TLR4
|
Asp299Gly
+896
(A→G) |
rs4986790 |
AA |
AG |
GG |
A |
G |
AA |
AG |
GG |
A |
G |
|
| 155 |
40 |
0 |
NA |
NA |
194 |
37 |
2 |
NA |
NA |
23 |
| 86 |
18 |
0 |
NA |
NA |
21 |
5 |
0 |
NA |
NA |
20 |
| 88 |
15 |
0 |
NA |
NA |
42 |
4 |
0 |
NA |
NA |
25 |
| 524 |
69 |
1 |
NA |
NA |
313 |
45 |
0 |
NA |
NA |
27 |
| 206 |
25 |
1 |
NA |
NA |
222 |
28 |
4 |
NA |
NA |
29 |
| 45 |
25 |
7 |
115 |
39 |
189 |
38 |
3 |
416 |
44 |
19 |
| 50 |
NA |
NA |
106 |
6 |
41 |
NA |
NA |
85 |
3 |
22 |
Thr399Ile
1196
(C→T) |
rs4986791 |
CC |
CT |
TT |
C |
T |
CC |
CT |
TT |
C |
T |
|
| 93 |
11 |
0 |
NA |
NA |
21 |
5 |
0 |
NA |
NA |
20 |
| 38 |
30 |
9 |
106 |
48 |
202 |
22 |
6 |
426 |
34 |
19 |
| 49 |
NA |
NA |
103 |
9 |
42 |
NA |
NA |
86 |
2 |
22 |
+3725
(G→C) |
rs11536889 |
GG |
GC |
CC |
G |
C |
GG |
GC |
CC |
G |
C |
|
| 627 |
474 |
90 |
NA |
NA |
199 |
162 |
40 |
NA |
NA |
15 |
| 279 |
51 |
9 |
609 |
69 |
182 |
44 |
4 |
408 |
52 |
28 |
| 110 |
62 |
10 |
282 |
82 |
50 |
36 |
3 |
136 |
42 |
13 |
| 37 |
NA |
NA |
93 |
19 |
26 |
NA |
NA |
70 |
18 |
22 |
| T→C |
rs10759932 |
TT |
TC |
CC |
T |
C |
TT |
TC |
CC |
T |
C |
|
| 153 |
37 |
14 |
NA |
NA |
139 |
53 |
4 |
NA |
NA |
16 |
| C→T |
rs1927914 |
CC |
CT |
TT |
C |
T |
CC |
CT |
TT |
C |
T |
|
| 0 |
8 |
69 |
8 |
146 |
0 |
150 |
80 |
150 |
310 |
19 |
TLR5
|
+1174
(C→T) |
rs5744168 |
CC |
CT |
TT |
C |
T |
CC |
CT |
TT |
C |
T |
|
| 206 |
19 |
0 |
NA |
NA |
239 |
16 |
0 |
NA |
NA |
29 |
TLR9 |
C→T |
rs352140 |
CC |
CT |
TT |
C |
T |
CC |
CT |
TT |
C |
T |
|
| 25 |
46 |
6 |
96 |
58 |
100 |
98 |
32 |
298 |
162 |
19 |
C→T |
rs34399053 |
CC |
CT |
TT |
C |
T |
CC |
CT |
TT |
C |
T |
|
| 26 |
51 |
0 |
103 |
51 |
79 |
150 |
1 |
308 |
152 |
19 |
C→T |
rs150459369 |
CC |
CT |
TT |
C |
T |
CC |
CT |
TT |
C |
T |
|
| 77 |
0 |
0 |
154 |
0 |
220 |
10 |
0 |
450 |
10 |
19 |
-1237
(T→C) |
rs5743836 |
TT |
TC |
CC |
T |
C |
TT |
TC |
CC |
T |
C |
|
| 77 |
40 |
0 |
NA |
NA |
35 |
15 |
1 |
NA |
NA |
24 |
TLR10
|
C→T |
rs10004195 |
AA |
AT |
TT |
A |
T |
AA |
AT |
TT |
A |
T |
|
| 498 |
712 |
276 |
NA |
NA |
308 |
493 |
207 |
NA |
NA |
12 |
| 59 |
10 |
135 |
NA |
NA |
22 |
123 |
51 |
NA |
NA |
16 |
| 24 |
18 |
15 |
NA |
NA |
9 |
11 |
8 |
NA |
NA |
18 |
Ile775Val |
rs4129009 |
TT |
TC |
CC |
T |
C |
TT |
TC |
CC |
T |
C |
|
| 507 |
715 |
275 |
NA |
NA |
351 |
490 |
177 |
NA |
NA |
12 |
TLR= Toll-like receptor; SNP= Single nucleotide polymorphism; ins= Insertion, del= Deletion; NA= Not available
Table 3. Association between polymorphisms in the TLRs genes, genotypes/alleles, and H. pylori infection
| Polymorphism |
Genotype/Allele |
Odds Ratio |
95% CI |
P-value |
I2
(%) |
Effect
model |
| TLR1 rs4833095 |
C vs. T |
NA |
NA |
- |
- |
- |
|
CC vs. TT |
1.605 |
(0.485-5.309) |
0.439 |
69.2 |
R |
|
CC vs. CT |
0.048 |
(0.002-1.247) |
0.068 |
98.4 |
R |
| TLR1 rs5743618 |
I vs. S |
NA |
NA |
- |
- |
- |
| |
II vs. SS |
1.277 |
(0.394-4.136) |
0.684 |
0.00 |
F |
| |
II vs. IS |
1.210 |
(0.354-4.142) |
0.304 |
0.00 |
F |
| TLR2 -196 to -174 ins>del |
ins vs. del |
0.876 |
(0.366-2.097) |
0.766 |
79 |
R |
| |
ins/ins vs. del/del |
1.028 |
(0.754-1.402) |
0.862 |
27 |
F |
| |
ins/ins vs. ins/del |
0.995 |
(0.821-1.208) |
0.963 |
19.2 |
F |
| TLR2 rs3804099 |
T vs. C |
NA |
NA |
- |
- |
- |
| |
TT vs. CC |
2.209 |
(1.283-3.804) |
0.004 |
0.00 |
F |
| |
TT vs. TC |
0.685 |
(0.397-1.179) |
0.172 |
0.00 |
F |
| TLR2 rs3804100 |
T vs. C |
NA |
NA |
- |
- |
- |
| |
TT vs. CC |
2.027 |
(0.597-6.878) |
0.257 |
0.00 |
F |
| |
TT vs. TC |
1.155 |
(0.723-1.846) |
0.547 |
0.00 |
F |
| TLR2 (+2251) |
G vs. A |
NA |
NA |
- |
- |
- |
| |
GG vs. AA |
- |
- |
- |
- |
- |
| |
GG vs. GA |
0.560 |
(0.102-3.087) |
0.506 |
0.00 |
F |
| TLR2 rs121917864 |
C vs. T |
0.980 |
(0.561-1.712) |
0.943 |
0.00 |
F |
| |
CC vs. TT |
NA |
NA |
- |
- |
- |
| |
CC vs. CT |
- |
- |
- |
- |
- |
| TLR2 rs5743708 |
G vs. A |
6.980 |
(0.856-56.901) |
0.070 |
0.00 |
F |
| |
GG vs. AA |
NA |
NA |
- |
- |
- |
| |
GG vs. GA |
NA |
NA |
- |
- |
- |
| TLR4 rs4986790 |
A vs. G |
2.987 |
(1.899-4.697) |
0.000 |
0.00 |
F |
| |
AA vs. GG |
1.234 |
(0.148-10.261) |
0.846 |
69.6 |
R |
| |
AA vs. AG |
1.293 |
(0.887-1.884) |
0.182 |
53.1 |
R |
| TLR4 rs4986791 |
C vs. T |
5.469 |
(3.432-8.713) |
0.000 |
0.00 |
F |
| |
CC vs. TT |
7.974 |
(2.682-23.706) |
0.000 |
0.00 |
F |
| |
CC vs. CT |
1.984 |
(0.144-27.401) |
0.609 |
93.6 |
R |
| TLR4 rs11536889 |
G vs. C |
0.895 |
(0.688-1.165) |
0.410 |
0.00 |
F |
| |
GG vs. CC |
0.813 |
(0.562-1.175) |
0.271 |
12.3 |
F |
| |
GG vs. GC |
0.872 |
(0.717-1.060) |
0.169 |
0.00 |
F |
| TLR4 rs10759932 |
T vs. C |
NA |
NA |
- |
- |
- |
| |
TT vs. CC |
3.180 |
(1.022-9.890) |
0.040 |
0.00 |
F |
| |
TT vs. TC |
0.634 |
(0.393-1.024) |
0.062 |
0.00 |
F |
| TLR4 rs1927914 |
C vs. T |
8.831 |
(4.222-18.470) |
0.000 |
0.00 |
F |
| |
CC vs. TT |
- |
- |
- |
- |
- |
| |
CC vs. CT |
- |
- |
- |
- |
- |
| TLR5 rs5744168 |
C vs. T |
NA |
NA |
- |
- |
- |
| |
CC vs. TT |
- |
- |
- |
- |
- |
| |
CC vs. CT |
1.378 |
(0.691-2.749) |
0.363 |
0.00 |
F |
| TLR9 rs352140 |
C vs. T |
1.111 |
(0.762-1.622) |
0.584 |
0.00 |
F |
| |
CC vs. TT |
0.750 |
(0.283-1.990) |
0.563 |
0.00 |
F |
| |
CC vs. CT |
1.878 |
(1.071-3.290) |
0.028 |
0.00 |
F |
| TLR9 rs34399053 |
C vs. T |
1.003 |
(0.681-1.479) |
0.987 |
0.00 |
F |
| |
CC vs. TT |
1.000 |
(0.040-25.296) |
1.000 |
0.00 |
F |
| |
CC vs. CT |
1.033 |
(0.599-1.782) |
0.907 |
0.00 |
F |
| TLR9 rs150459369 |
C vs. T |
0.139 |
(0.008-2.383) |
0.173 |
0.00 |
F |
| |
CC vs. TT |
- |
- |
- |
- |
- |
| |
CC vs. CT |
0.135 |
(0.008-2.339) |
0.169 |
0.00 |
F |
| TLR9 rs5743836 |
T vs. C |
NA |
NA |
- |
- |
- |
| |
TT vs. CC |
0.153 |
(0.006-3.841) |
0.253 |
0.00 |
F |
| |
TT vs. TC |
1.212 |
(0.593-2.479) |
0.598 |
0.00 |
F |
| TLR10 rs10004195 |
A vs. T |
NA |
NA |
- |
- |
- |
| |
AA vs. TT |
0.839 |
(0.680-1.036) |
0.102 |
0.00 |
F |
| |
AA vs. AT |
0.258 |
(0.029-2.288) |
0.224 |
96.8 |
R |
| TLR10 rs4129009 |
T vs. C |
NA |
NA |
- |
- |
- |
| |
TT vs. CC |
1.076 |
(0.852-1.358) |
0.539 |
0.00 |
F |
|
TT vs. TC |
1.010 |
(0.845-1.207) |
0.911 |
0.00 |
F |
Bold= Statistically significant result; R= Random-effect model; F= Fixed-effect model; CI= Confidence interval; NA= Not available
Discussion
Genetic polymorphisms in TLRs, as key members of the innate immune system, mediate recognition of
H. pylori and are assumed to play an important role in susceptibility to
H. pylori infections [30]. However, there have been many conflicting reports concerning TLRs polymorphisms and their possible role in susceptibility to
H. pylori infections. The current study is the first comprehensive systematic review and meta-analysis on the evaluation of association between TLR1, 2, 4, 5, 9 and 10 polymorphisms with of
H. pylori infection risk. TLR1 is a PRR which forms heterodimer with TLR2 and recognizes lipoprotein/lipopeptides of
H. pylori [31]. Recently, several studies have shown that individuals carrying TLR1 rs4833095 CT and TT genotypes and T allele as well as TLR1 rs5743618 SS genotype significantly decrease the risks of
H. pylori infection and
H. pylori-related diseases [12, 26]. In the same manner, individuals carrying CC or TT homozygous genotypes are at increased risks [16-18]. However, in this study, the genotypic model failed to show any significant association between TLR1 rs4833095 and
rs5743618 polymorphisms and the susceptibility to
H. pylori infections (Table 3). This difference may be due to the differences in ethnicities, assessment methods, population size and age.
Another member of the PRRs is TLR2 which is implicated in the identification of Gram-positive bacteria, mycobacteria, spirochetes, viruses, hepatitis C and B viruses, herpes simplex and cytomegalovirus, and fungi [31]. In the case of
H. pylori, SNPs in genes that encode TLR2 are associated with
H. pylori infection and
H. pylori-related diseases.
Our systematic review of included articles revealed that TLR2
-196 to 174del polymorphism is not significantly associated with
H. pylori infection in the Japanese population [14]. However, increased risk of
H. pylori-related diseases in
H. pylori-infected individuals was observed in Chinese and Iranian populations carrying the above-mentioned polymorphism [13, 21]. In this meta-analysis, we could not detect any association between TLR2
-196 to 174del polymorphism and susceptibility to
H. pylori infections (Table 3). This difference may arise from variations in ethnicities. In the studies conducted in the Thai population, no significant association was discerned between TLR2 rs3804099 and
rs3804100 polymorphisms and susceptibility to
H. pylori infections [16]. However, we observed that individuals with the CC mutant homozygote genotype for TLR2 rs3804099 bear a significantly increased risk of
H. pylori infection (odds ratio = 2.209, 95% CI: 1.283-3.804) (Table 3). In the Brazilian population, TLR2 (+2251) failed to project a significant association with susceptibility to
H. pylori infections while in the Iranian population TLR2 rs121917864 and
rs5743708 proved significant associations with
H. pylori infection [21, 29]. Our findings suggest that the TLR2 rs5743708 A allele mutant was significantly associated and susceptibility to
H. pylori infections (odds ratio = 6.980, 95% CI: 0.856-56.901) (Table 3).
TLR4 is located on the surface of immune cells such as monocytes, mast cells and neutrophils and similar to TLR2, recognizes a wide number of endogenous ligands released during cellular stress and necrosis, and preserved microbial structures of Gram-negative bacteria [31]. TLR4 expression increases during
H. pylori infection on gastric epithelial cells and is the LPS receptor [31]. In the current meta-analysis, five SNPs were found in the human TLR4 gene in
H. pylori positive subjects.
Whilst several studies with different population have reported that the TLR4 polymorphisms
rs4986790,
rs4986791,
rs11536889,
rs10759932 and
rs1927914 are either associated or not associated with the risk of
H. pylori infection (Table 2), our study offers a significant association only with the TLR4 rs4986790 G mutant allele (odds ratio = 2.987, 95% CI: 1.899-4.697), TLR4 rs4986791 TT mutant homozygote genotype (odds ratio = 7.974, 95% CI: 2.682-23.706) and T mutant allele (odds ratio = 5.469, 95% CI: 13.432-8.713), TLR4 rs10759932 CC mutant genotype (odds ratio = 3.180, 95% CI: 1.022-9.890) and TLR4 rs1927914 T mutant allele (odds ratio = 8.831, 95% CI: 4.222-18.470) and susceptibility to
H. pylori infections (Table 3 and Fig. 2).
Monomeric bacterial flagellin has the ability to stimulate innate immune responses mediated by TLR5 which is located on surface epithelial, and membrane of immune cells such as NK cells, monocytes and myeloid dendritic cells [31]. There are controversial studies on recognizing
H. pylori flagellin by TLR5 and the association of TLR5 polymorphism and susceptibility to
H. pylori infection [29]. In the current meta-analysis, there was no evidence linking TLR5 polymorphism and susceptibility to
H. pylori infection.
The plasmacytoid dendritic cells, B cells and NK cells express TLR9 which is involved in stimulating innate immune responses via detection of bacterial and viral unmethylated CpG DNA [31]. It has also been suggested that TLR9 rs352140 and
s5743836 poly-morphisms play a role in the susceptibility to
H. pylori infection [19, 24]. Our results also demonstrated that TLR9 rs352140 CT mutant genotype conferred a significantly increased risk of
H. pylori infections (odds ratio = 1.878, 95% CI: 1.071-3.290) (Table 3 and Fig. 2).
TLR10 is an anti-inflammatory PRR which is expressed in humans [12, 31]. Tang et al. [10], Simawaranon et al. [16], and Ram et al. [18] reported that TLR10 rs10004195 is associated with susceptibility to
H. pylori infection in the Chinese, Thai and Malaysian population. However, no association was found between TLR10 polymorphisms and susceptibility to
H. pylori infections in our study (Table 3).
TLR11, TLR12 and TLR13 are expressed in mice but we could not find any study on the association of TLR3, 6-8 and 11-13 polymorphisms and susceptibility to
H. pylori infections.
Conclusion
This meta-analysis indicated that TLR2 rs3804099, TLR4 rs4986790, TLR4 rs4986791, TLR4 rs10759932, TLR4 rs1927914 and TLR9 rs352140 are associated and increased susceptibility to
H. pylori infections. Possible reasons for discrepant findings in different studies may be variances in population size, age and sex, ethnicity and race, methods of diagnosis of infection and genotyping, and differences in the prevalence of
H. pylori infection. The results of this meta-analysis led to the widely accepted conclusion that
H. pylori infections are associated with TLRs genetic variations. In conclusion, evidences support the important role of TLRs in
H. pylori infection as these receptors of the innate immune system have been shown to recognize diverse components of
H. pylori, the major risk factor for gastric cancer. Given that host genetic variability in the TLRs are known to be associated with an increased risk of
H. pylori infection, this knowledge has the potential to allow better prevention of
H. pylori infection and subsequently gastric cancer through selective treatment and surveillance of individuals harboring high risk genetic profiles.
Conflict of Interest
None.
Acknowledgments
We would like to thank all the individuals who helped in this study, especially Sahar Sabour, Zahra Hosseinali, and other colleagues in the department of microbiology, Ardabil University of Medical Sciences.
References
- Khademi F, Faghri J, Moghim S, Esfahani BN, Fazeli H, Poursina F, et al. The study of mutation in 23S rRNA resistance gene of Helicobacter pylori to clarithromycin in patients with gastrointestinal disorders in Isfahan, Iran. Adv Biomed Res. 2014; 3(98): 1-4.
- Khademi F, Poursina F, Hosseini E, Akbari M, Safaei HG. Helicobacter pylori in Iran: A systematic review on the antibiotic resistance. Iran J Basic Med Sci. 2015; 18(1): 2-7.
- Yousefi-Avarvand A, Vaez H, Tafaghodi M, Sahebkar AH, Arzanlou M, Khademi F. Antibiotic resistance of helicobacter pylori in Iranian children: A systematic review and meta-analysis. Microb Drug Resist. 2018; 24(7): 980-86.
- Fan Yf, Wu YM, Liu H, Yu Y, Jiang Yy, Xue Yz, et al. TLR4 polymorphisms associated with developing gastric pre-cancer lesions in a Chinese Han population. Hum Immunol. 2014; 75(2): 176-81.
- Kusters JG, van Vliet AH, Kuipers EJ. Pathogenesis of Helicobacter pylori infection. Clin Microbiol Rev. 2006; 19(3): 449-90.
- Achyut B, Ghoshal UC, Moorchung N, Mittal B. Association of Toll-like receptor–4 (Asp299Gly and Thr399Ileu) gene polymorphisms with gastritis and precancerous lesions. Hum Immunol. 2007; 68(11): 901-907.
- Torres J, Pérez-Rodríguez M, Camorlinga-Ponce M, Luna LF, Abdo-Francis JM, Lazcano E, et al. TLR4 single-nucleotide polymorphisms alter mucosal cytokine and chemokine patterns in Mexican patients with Helicobacter pylori-associated gastroduodenal diseases. Clin Immunol. 2008; 129(2): 333-40.
- Kutikhin AG. Association of polymorphisms in TLR genes and in genes of the Toll-like receptor signaling pathway with cancer risk. Hum Immunol. 2011; 72(11): 1095-116.
- Khademi F, Derakhshan M, Sadeghi R. The role of toll-like receptor gene polymorphisms in tuberculosis susceptibility: A systematic review and meta-analysis. Rev Clin Med. 2016; 3(4): 133-40.
- Chang Z. Important aspects of Toll-like receptors, ligands and their signaling pathways. Inflamm Res. 2010; 59(10): 791-808.
- Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ: British Medical Journal. 2003; 327(7414): 557-65.
- Tang Fb, Li Zx, Wang Ym, Zhang L, Ma Jl, Zhou T, et al. Toll-like receptor 1 and 10 polymorphisms, Helicobacter pylori suspec-tibility and risk of gastric lesions in a high-risk Chinese population. Infect Genet Evol. 2015; 31(1): 263-69.
- Castaño-Rodríguez N, Kaakoush NO, Goh K-L, Fock KM, Mitchell HM. The role of TLR2, TLR4 and CD14 genetic polymorphisms in gastric carcinogenesis: a case-control study and meta-analysis. PLoS One 2013; 8(4): 60327.
- Hishida A, Matsuo K, Goto Y, Naito M, Wakai K, Tajima K, et al. No associations of Toll-like receptor 2 (TLR2)-196 to-174del polymorphism with the risk of Helicobacter pylori seropositivity, gastric atrophy, and gastric cancer in Japanese. Gastric Cancer 2010; 13(4): 251-57.
- Hishida A, Matsuo K, Goto Y, Mitsuda Y, Hiraki A, Naito M, et al. Toll‐like receptor 4+ 3725 G/C polymorphism, helicobacter pylori seropositivity, and the risk of gastric atrophy and gastric cancer in Japanese. Helicobacter 2009; 14(1): 47-53.
- Simawaranon T, Wattanawongdon W, Tongtawee T. Toll-like receptors are associated with helicobacter pylori infection and gastric mucosa pathology. Jundishapur J Microbiol. 2017; 10(12): 1-9.
- Tongtawee T, Bartpho T, Kaewpitoon S, Kaewpitoon N, Dechsukhum C, Leeanansaksiri W, et al. TLR1 polymorphism associations with gastric mucosa morphologic patterns on magnifying NBI Endoscopy: a prospective crosssectional study. Asian Pac J Cancer Prev. 2016; 17(1): 3391-394.
- Ram MR, Goh KL, Leow AHR, Poh BH, Loke MF, Harrison R, et al. Polymorphisms at locus 4p14 of Toll-like receptors TLR-1 and TLR-10 confer susceptibility to gastric carcinoma in Helicobacter pylori infection. PloS one 2015; 10(11): 141865.
- Loganathan R, Nazeer M, Goda V, Devaraju P, Ali M, Karunakaran P, et al. Genetic variants of TLR4 and TLR9 are risk factors for chronic Helicobacter pylori infection in South Indian Tamils. Hum Immunol. 2017; 78(2): 216-20.
- Qadri Q, Rasool R, Afroze D, Naqash S, Gulzar G, Yousuf A, et al. Study of TLR4 and IL-8 gene polymorphisms in H. pylori-induced inflammation in gastric cancer in an ethnic Kashmiri population. Immunol Invest. 2014; 43(4): 324-36.
- Habibzadeh M, Tourani M, Shokri-Shirvani J, Mostafazadeh A, Khafri S, Nouri HR. TLR2 Arg677Trp but not TLR2 -196 to -174 ins/del and Arg753Gln polymorphism alter the risk of peptic ulcer in north of Iran. J Chinese Med Associ. 2017; 5(4):1-6.
- Tourani M, Habibzadeh M, Shokri‐Shirvani J, Teymournejad O, Mostafazadeh A, Khafri S, et al. Association of Helicobacter pylori infection with Toll‐like receptor‐4 Thr399Ile polymorphism increased the risk of peptic ulcer development in North of Iran. APMIS. 2018; 126(1): 76-84.
- Bagheri N, Azadegan-Dehkordi F, Sanei H, Taghikhani A, Rahimian G, Salimzadeh L, et al. Associations of a TLR4 single-nucleotide polymorphism with Helicobacter pylori associated gastric diseases in iranian patients. Clin Res Hepatol Gastroenterol. 2014; 38(3): 366-71.
- Ng MTH, Van't Hof R, Crockett JC, Hope ME, Berry S, Thomson J, et al. Increase in NF-κB binding affinity of the variant C allele of the Toll-like receptor 9− 1237T/C polymorphism is associated with Helicobacter pylori-induced gastric disease. Infect Immun. 2010; 78(3): 1345-352.
- Hold GL, Rabkin CS, Chow WH, Smith MG, Gammon MD, Risch HA, et al. A functional polymorphism of toll-like receptor 4 gene increases risk of gastric carcinoma and its precursors. Gastroenterology 2007; 132(3): 905-12.
- Yang CA, Scheibenbogen C, Bauer S, Kleinle C, Wex T, Bornschein J, et al. A frequent toll‐like receptor 1 gene polymorphism affects NK‐and T‐cell IFN‐γ production and is associated with helicobacter pylori‐induced gastric disease. Helicobacter 2013; 18(1): 13-21.
- Hellmig S, Fischbach W, Goebeler-Kolve ME, Fölsch UR, Hampe J, Schreiber S. Association study of a functional Toll-like receptor 4 polymorphism with susceptibility to gastric mucosa-associated lymphoid tissue lymphoma. Leuk Lymphoma. 2005; 46(6): 869-72.
- Kupcinskas J, Wex T, Bornschein J, Selgrad M, Leja M, Juozaityte E, et al. Lack of association between gene polymorphisms of Angiotensin converting enzyme, Nod-like receptor 1, Toll-like receptor 4, FAS/FASL and the presence of Helicobacter pylori-induced premalignant gastric lesions and gastric cancer in Caucasians. BMC Med Genet. 2011; 12(1): 112.
- Moura SB, Almeida LR, Guerra JB, Rocha GA, Rocha AMC, Melo FF, et al. Toll-like receptor (TLR2, TLR4 and TLR5) gene polymorphisms and Helicobacter pylori infection in children with and without duodenal ulcer. Microbes Infect. 2008; 10(14-15): 1477-483.
- El-Omar E, Ng M, Hold G. Polymorphisms in Toll-like receptor genes and risk of cancer. Oncogene. 2008; 27(2): 244.
- Testro AG, Visvanathan K. Toll‐like receptors and their role in gastrointestinal disease. J Gastroenterol Hepatol. 2009; 24(6): 943-54.










































 , Amirhossein Sahebkar
, Amirhossein Sahebkar 

 , Asad Mohammadi
, Asad Mohammadi 

 , Mohsen Arzanlou
, Mohsen Arzanlou 

 , Arshid Yousefi-Avarvand
, Arshid Yousefi-Avarvand 

 , Farzad Khademi *
, Farzad Khademi * 




